Example 1 - Continuous and categorical predictors
using Metida, CSV, DataFrames, CategoricalArrays, MixedModels;
rds = CSV.File(joinpath(dirname(pathof(Metida)), "..", "test", "csv", "1fptime.csv"); types = [String, String, Float64, Float64]) |> DataFrame
rds2 = CSV.File(joinpath(dirname(pathof(Metida)), "..", "test", "csv", "ftdf3.csv"); types = [String, Float64, Float64, String, String, String, String, String, Float64]) |> DataFrame
devday = CSV.File(joinpath(dirname(pathof(Metida)), "..", "test", "csv", "devday.csv"); types = [Float64, String, String, String ]) |> DataFrame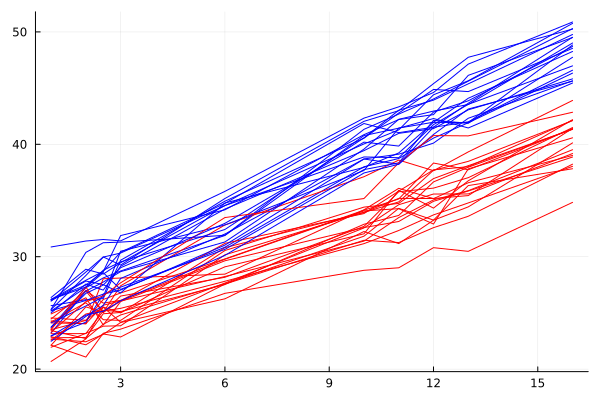
Metida result:
lmm = LMM(@formula(response ~1 + factor*time), rds;
random = VarEffect(@covstr(1 + time|subject&factor), CSH),
)
fit!(lmm)Linear Mixed Model: response ~ 1 + factor + time + factor & time
Random 1:
Model: :(1 + time)|:(subject & factor)
Type: CSH (3), Subjects: 40
Repeated:
Residual only
Blocks: 40, Maximum block size: 10
Status: converged (No Errors)
-2 logREML: 1300.18 BIC: 1324.11
Fixed-effects parameters:
──────────────────────────────────────────────────────────
Coef. Std. Error z Pr(>|z|)
──────────────────────────────────────────────────────────
(Intercept) 22.1331 0.304339 72.73 <1e-99
factor: 1.0 2.00049 0.430401 4.65 <1e-05
time 1.11853 0.0264038 42.36 <1e-99
factor: 1.0 & time 0.404971 0.0373406 10.85 <1e-26
──────────────────────────────────────────────────────────
Variance components:
θ vector: [1.2401, 0.101122, -0.0427208, 0.985061]
Random 1 σ² (Intercept) var 1.53785
Random 1 σ² time var 0.0102257
Random 1 ρ rho -0.0427208
Residual σ² var 0.970345
MixedModels result:
fm = @formula(response ~ 1 + factor*time + (1 + time|subject&factor))
mm = fit(MixedModel, fm, rds, REML=true)
Minimizing 2 Time: 0:00:00 ( 0.14 s/it)
objective: 1426.9231858044589
Minimizing 66 Time: 0:00:00 ( 5.13 ms/it)
Linear mixed model fit by REML
response ~ 1 + factor + time + factor & time + (1 + time | subject & factor)
REML criterion at convergence: 1300.18075981887
Variance components:
Column Variance Std.Dev. Corr.
subject & factor (Intercept) 1.537868 1.240108
time 0.010226 0.101122 -0.04
Residual 0.970345 0.985061
Number of obs: 400; levels of grouping factors: 40
Fixed-effects parameters:
──────────────────────────────────────────────────────────
Coef. Std. Error z Pr(>|z|)
──────────────────────────────────────────────────────────
(Intercept) 22.1331 0.30434 72.72 <1e-99
factor: 1.0 2.00049 0.430402 4.65 <1e-05
time 1.11853 0.0264037 42.36 <1e-99
factor: 1.0 & time 0.404971 0.0373405 10.85 <1e-26
──────────────────────────────────────────────────────────Example 2 - Two random factors (Penicillin data)
Metida:
df = CSV.File(joinpath(dirname(pathof(Metida)), "..", "test", "csv", "Penicillin.csv"); types = [String, Float64, String, String]) |> DataFrame
df.diameter = float.(df.diameter)
lmm = LMM(@formula(diameter ~ 1), df;
random = [VarEffect(@covstr(1|plate), SI), VarEffect(@covstr(1|sample), SI)]
)
fit!(lmm)Linear Mixed Model: diameter ~ 1
Random 1:
Model: 1|plate
Type: SI (1), Subjects: 24
Random 2:
Model: 1|sample
Type: SI (1), Subjects: 6
Repeated:
Residual only
Blocks: 1, Maximum block size: 144
Status: converged (No Errors)
-2 logREML: 330.861 BIC: 345.749
Fixed-effects parameters:
─────────────────────────────────────────────────
Coef. Std. Error z Pr(>|z|)
─────────────────────────────────────────────────
(Intercept) 22.9722 0.808573 28.41 <1e-99
─────────────────────────────────────────────────
Variance components:
θ vector: [0.846704, 1.93156, 0.549923]
Random 1 σ² var 0.716908
Random 2 σ² var 3.73092
Residual σ² var 0.302415
MixedModels:
fm2 = @formula(diameter ~ 1 + (1|plate) + (1|sample))
mm = fit(MixedModel, fm2, df, REML=true)
Minimizing 2 Time: 0:00:00 ( 0.26 s/it)
objective: 366.5302937135882
Minimizing 46 Time: 0:00:00 (11.22 ms/it)
Linear mixed model fit by REML
diameter ~ 1 + (1 | plate) + (1 | sample)
REML criterion at convergence: 330.86058899096145
Variance components:
Column Variance Std.Dev.
plate (Intercept) 0.716908 0.846704
sample (Intercept) 3.730907 1.931556
Residual 0.302415 0.549923
Number of obs: 144; levels of grouping factors: 24, 6
Fixed-effects parameters:
─────────────────────────────────────────────────
Coef. Std. Error z Pr(>|z|)
─────────────────────────────────────────────────
(Intercept) 22.9722 0.808572 28.41 <1e-99
─────────────────────────────────────────────────Example 3 - Repeated ARMA/AR/ARH
rds = CSV.File(joinpath(dirname(pathof(Metida)), "..", "test", "csv", "1freparma.csv"); types = [String, String, Float64, Float64]) |> DataFrame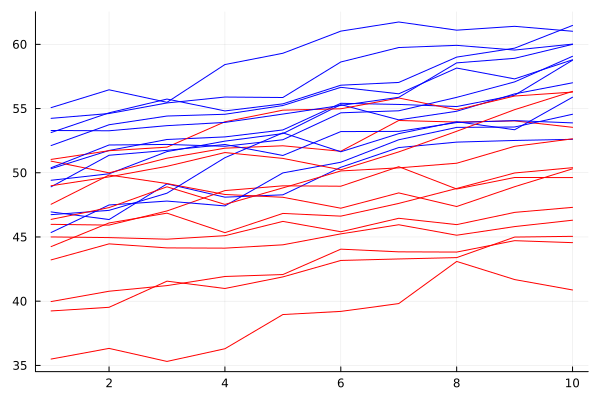
ARMA:
lmm = LMM(@formula(response ~ 1 + factor*time), rds;
random = VarEffect(@covstr(factor|subject&factor), DIAG),
repeated = VarEffect(@covstr(1|subject&factor), ARMA),
)
fit!(lmm)Linear Mixed Model: response ~ 1 + factor + time + factor & time
Random 1:
Model: factor|:(subject & factor)
Type: DIAG (2), Subjects: 24
Repeated:
Model: 1|:(subject & factor)
Type: ARMA (3)
Blocks: 24, Maximum block size: 10
Not fitted.
AR:
lmm = Metida.LMM(@formula(response ~ 1 + factor*time), rds;
random = VarEffect(@covstr(factor|subject&factor), DIAG),
repeated = VarEffect(@covstr(1|subject&factor), AR),
)
fit!(lmm)Linear Mixed Model: response ~ 1 + factor + time + factor & time
Random 1:
Model: factor|:(subject & factor)
Type: DIAG (2), Subjects: 24
Repeated:
Model: 1|:(subject & factor)
Type: AR (2)
Blocks: 24, Maximum block size: 10
Status: converged (No Errors)
-2 logREML: 710.096 BIC: 731.952
Fixed-effects parameters:
──────────────────────────────────────────────────────────
Coef. Std. Error z Pr(>|z|)
──────────────────────────────────────────────────────────
(Intercept) 44.3915 1.34686 32.96 <1e-99
factor: 1.0 5.39757 1.60271 3.37 0.0008
time 0.508075 0.0456524 11.13 <1e-28
factor: 1.0 & time 0.290317 0.0645622 4.50 <1e-05
──────────────────────────────────────────────────────────
Variance components:
θ vector: [4.54797, 2.82342, 1.05771, 0.576979]
Random 1 σ² factor: 0.0 var 20.684
Random 1 σ² factor: 1.0 var 7.9717
Residual σ² var 1.11875
Residual ρ rho 0.576979
ARH:
lmm = Metida.LMM(@formula(response ~ 1 + factor*time), rds;
random = VarEffect(@covstr(factor|subject&factor), DIAG),
repeated = VarEffect(@covstr(1|subject&factor), ARH),
)
fit!(lmm)Linear Mixed Model: response ~ 1 + factor + time + factor & time
Random 1:
Model: factor|:(subject & factor)
Type: DIAG (2), Subjects: 24
Repeated:
Model: 1|:(subject & factor)
Type: ARH (2)
Blocks: 24, Maximum block size: 10
Status: converged (No Errors)
-2 logREML: 710.096 BIC: 731.952
Fixed-effects parameters:
──────────────────────────────────────────────────────────
Coef. Std. Error z Pr(>|z|)
──────────────────────────────────────────────────────────
(Intercept) 44.3915 1.34686 32.96 <1e-99
factor: 1.0 5.39757 1.60271 3.37 0.0008
time 0.508075 0.0456524 11.13 <1e-28
factor: 1.0 & time 0.290317 0.0645622 4.50 <1e-05
──────────────────────────────────────────────────────────
Variance components:
θ vector: [4.54797, 2.82342, 1.05771, 0.576979]
Random 1 σ² factor: 0.0 var 20.684
Random 1 σ² factor: 1.0 var 7.9717
Residual σ² (Intercept) var 1.11875
Residual ρ rho 0.576979
Example 4 - SAS relation
Model 1
df0 = CSV.File(joinpath(dirname(pathof(Metida)), "..", "test", "csv", "df0.csv")) |> DataFrame
lmm = LMM(@formula(var ~ sequence + period + formulation), df0;
random = VarEffect(@covstr(formulation|subject), CSH),
repeated = VarEffect(@covstr(formulation|subject), DIAG),
)
fit!(lmm)SAS code:
PROC MIXED data=df0;
CLASSES subject sequence period formulation;
MODEL var = sequence period formulation/ DDFM=SATTERTH s;
RANDOM formulation/TYPE=CSH SUB=subject G V;
REPEATED/GRP=formulation SUB=subject R;
RUN;Model 2
lmm = LMM(
@formula(var ~ sequence + period + formulation), df0;
random = VarEffect(@covstr(formulation|subject), SI),
repeated = VarEffect(@covstr(formulation|subject), DIAG),
)
fit!(lmm)SAS code:
PROC MIXED data=df0;
CLASSES subject sequence period formulation;
MODEL var = sequence period formulation/ DDFM=SATTERTH s;
RANDOM formulation/TYPE=VC SUB=subject G V;
REPEATED/GRP=formulation SUB=subject R;
RUN;Model 3
lmm = LMM(@formula(var ~ sequence + period + formulation), df0;
random = VarEffect(@covstr(subject|1), SI)
)
fit!(lmm)SAS code:
PROC MIXED data=df0;
CLASSES subject sequence period formulation;
MODEL var = sequence period formulation/ DDFM=SATTERTH s;
RANDOM subject/TYPE=VC G V;
RUN;Example 5 - Working with Effects.jl
using Effects, StatsModels
lmm = LMM(@formula(var ~ sequence + period + formulation), df0;
random = VarEffect(@covstr(subject|1), SI)
)
fit!(lmm)
table_model = StatsModels.TableRegressionModel(lmm, lmm.mf, lmm.mm)
emmeans(tm)
effects(Dict(:period => ["1", "2", "3", "4"]), tm)Unstructured covariance
Unstructured covariance example.
Metida result:
lmm = Metida.LMM(@formula(response~factor), rds2;
random = Metida.VarEffect(Metida.@covstr(r1|subject), UN),
)
Metida.fit!(lmm)Linear Mixed Model: response ~ factor
Random 1:
Model: r1|subject
Type: UN (6), Subjects: 24
Repeated:
Residual only
Blocks: 24, Maximum block size: 7
Status: converged (No Errors)
-2 logREML: 705.382 BIC: 741.166
Fixed-effects parameters:
──────────────────────────────────────────────────
Coef. Std. Error z Pr(>|z|)
──────────────────────────────────────────────────
(Intercept) 40.1983 0.791652 50.78 <1e-99
factor -11.0271 1.11956 -9.85 <1e-22
──────────────────────────────────────────────────
Variance components:
θ vector: [2.91848, 3.56669, 4.59709, 0.458287, 0.463127, 0.575315, 1.00823]
Random 1 σ² r1: A var 8.51753
Random 1 σ² r1: B var 12.7213
Random 1 σ² r1: C var 21.1332
Random 1 ρ: r1: A × r1: B rho 0.458287
Random 1 ρ: r1: A × r1: C rho 0.463127
Random 1 ρ: r1: B × r1: C rho 0.575315
Residual σ² var 1.01653
MixedModels result:
mm = fit(MixedModel, @formula(response ~ factor+ (0+r1|subject)), rds2, REML = true)Linear mixed model fit by REML
response ~ 1 + factor + (0 + r1 | subject)
REML criterion at convergence: 705.3822332332761
Variance components:
ColumnVariance Std.Dev. Corr.
subject r1: A 8.51744 2.91847
r1: B 12.72007 3.56652 +0.46
r1: C 21.13152 4.59690 +0.46 +0.58
Residual 1.01654 1.00824
Number of obs: 168; levels of grouping factors: 24
Fixed-effects parameters:
──────────────────────────────────────────────────
Coef. Std. Error z Pr(>|z|)
──────────────────────────────────────────────────
(Intercept) 40.1982 0.791621 50.78 <1e-99
factor -11.0268 1.11952 -9.85 <1e-22
──────────────────────────────────────────────────Augmented covariance (Experimental)
Covariance modificator ACOV() can be used as second repeated effect. In this case covariance calculated with existed matrix, that was build at previous step. For example, addition ACOV(AR) to DIAG structure is the same as ARH if same blocking factor used.
lmm1 = Metida.LMM(@formula(response ~ 1), rds2;
repeated = [Metida.VarEffect(Metida.@covstr(r1|subject), Metida.DIAG), Metida.VarEffect(Metida.@covstr(1|subject), Metida.ACOV(Metida.AR))]
)
Metida.fit!(lmm1)Linear Mixed Model: response ~ 1
Random 1:
No
Repeated:
Model: r1|subject
Type: DIAG (3)
Model: 1|subject
Type: ACOV(AR) (1)
Blocks: 24, Maximum block size: 7
Status: converged (No Errors)
-2 logREML: 821.402 BIC: 841.874
Fixed-effects parameters:
─────────────────────────────────────────────────
Coef. Std. Error z Pr(>|z|)
─────────────────────────────────────────────────
(Intercept) 34.5061 0.961804 35.88 <1e-99
─────────────────────────────────────────────────
Variance components:
θ vector: [6.44278, 5.40009, 7.00676, 0.919938]
Residual 1 σ² r1: A var 41.5094
Residual 1 σ² r1: B var 29.161
Residual 1 σ² r1: C var 49.0947
Residual 2 ρ rho 0.919938
lmm2 = Metida.LMM(@formula(response ~ 1), rds2;
repeated = [Metida.VarEffect(Metida.@covstr(r1|subject), Metida.ARH)]
)
Metida.fit!(lmm2)Linear Mixed Model: response ~ 1
Random 1:
No
Repeated:
Model: r1|subject
Type: ARH (4)
Blocks: 24, Maximum block size: 7
Status: converged (No Errors)
-2 logREML: 821.402 BIC: 841.874
Fixed-effects parameters:
─────────────────────────────────────────────────
Coef. Std. Error z Pr(>|z|)
─────────────────────────────────────────────────
(Intercept) 34.5061 0.961804 35.88 <1e-99
─────────────────────────────────────────────────
Variance components:
θ vector: [6.44278, 5.40009, 7.00676, 0.919938]
Residual σ² r1: A var 41.5094
Residual σ² r1: B var 29.161
Residual σ² r1: C var 49.0947
Residual ρ rho 0.919938
R-part of variance-covariance matrix:
Metida.rmatrix(lmm1, 1)7×7 LinearAlgebra.Symmetric{Float64, Matrix{Float64}}:
41.5094 38.1861 35.1289 27.0863 24.9178 29.7429 27.3616
38.1861 41.5094 38.1861 29.4437 27.0863 32.3314 29.7429
35.1289 38.1861 41.5094 32.0061 29.4437 35.1452 32.3314
27.0863 29.4437 32.0061 29.161 26.8263 32.021 29.4574
24.9178 27.0863 29.4437 26.8263 29.161 34.8078 32.021
29.7429 32.3314 35.1452 32.021 34.8078 49.0946 45.164
27.3616 29.7429 32.3314 29.4574 32.021 45.164 49.0946If nested blocking factor used - covariance modification applyed only within that blocks (R-part of variance-covariance matrix is the same):
lmm = Metida.LMM(@formula(response ~ 1), rds2;
repeated = [Metida.VarEffect(Metida.@covstr(r1|subject), Metida.DIAG), Metida.VarEffect(Metida.@covstr(1|subject), Metida.ACOV(Metida.AR))]
)
Metida.fit!(lmm)
Metida.rmatrix(lmm, 1)7×7 LinearAlgebra.Symmetric{Float64, Matrix{Float64}}:
41.5094 38.1861 35.1289 27.0863 24.9178 29.7429 27.3616
38.1861 41.5094 38.1861 29.4437 27.0863 32.3314 29.7429
35.1289 38.1861 41.5094 32.0061 29.4437 35.1452 32.3314
27.0863 29.4437 32.0061 29.161 26.8263 32.021 29.4574
24.9178 27.0863 29.4437 26.8263 29.161 34.8078 32.021
29.7429 32.3314 35.1452 32.021 34.8078 49.0946 45.164
27.3616 29.7429 32.3314 29.4574 32.021 45.164 49.0946For nested models covariance structure can be expanded as follows:
- the first layer describes unstructured the device-device covariance;
- the second layer adds the time covariance for each device
lmm = Metida.LMM(@formula(resp ~ 0 + device), devday;
repeated = [Metida.VarEffect(Metida.@covstr(device|subj&day), Metida.UN),
Metida.VarEffect(Metida.@covstr(1|subj&device), Metida.ACOV(Metida.AR))]
)
Metida.fit!(lmm)Linear Mixed Model: resp ~ 0 + device
Random 1:
No
Repeated:
Model: device|:(subj & day)
Type: UN (10)
Model: 1|:(subj & device)
Type: ACOV(AR) (1)
Blocks: 25, Maximum block size: 8
Status: converged (No Errors)
-2 logREML: 903.058 BIC: 961.284
Fixed-effects parameters:
───────────────────────────────────────────────
Coef. Std. Error z Pr(>|z|)
───────────────────────────────────────────────
device: A 6.66689 0.856988 7.78 <1e-14
device: B 2.04241 0.322 6.34 <1e-09
device: C 1.26148 0.48472 2.60 0.0093
device: D 0.940181 0.634722 1.48 0.1385
───────────────────────────────────────────────
Variance components:
θ vector: [4.81355, 1.80862, 2.72258, 3.56512, 0.0857122, 0.137169, 0.347301, 0.296275, -0.0354115, 0.0300272, 0.584853]
Residual 1 σ² device: A var 23.1703
Residual 1 σ² device: B var 3.27111
Residual 1 σ² device: C var 7.41244
Residual 1 σ² device: D var 12.7101
Residual 1 ρ: device: A × device: B rho 0.0857122
Residual 1 ρ: device: A × device: C rho 0.137169
Residual 1 ρ: device: A × device: D rho 0.347301
Residual 1 ρ: device: B × device: C rho 0.296275
Residual 1 ρ: device: B × device: D rho -0.0354115
Residual 1 ρ: device: C × device: D rho 0.0300272
Residual 2 ρ rho 0.584853
R-matrix:
Metida.rmatrix(lmm, 1)8×8 LinearAlgebra.Symmetric{Float64, Matrix{Float64}}:
12.7101 7.43353 5.95998 … 0.0 -0.228331 0.0
7.43353 12.7101 0.0 0.291454 0.0 -0.228331
5.95998 0.0 23.1702 0.0 0.746199 0.0
0.0 5.95998 13.5512 1.79763 0.0 0.746199
0.291454 0.0 1.79763 4.3352 1.45889 0.0
0.0 0.291454 0.0 … 7.41246 0.0 1.45889
-0.228331 0.0 0.746199 0.0 3.2711 1.91311
0.0 -0.228331 0.0 1.45889 1.91311 3.2711